Gadolinium »
PDB 1b9x-4l12 »
3cqr »
Gadolinium in PDB 3cqr: Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5
Enzymatic activity of Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5
All present enzymatic activity of Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5:
1.10.99.3;
1.10.99.3;
Protein crystallography data
The structure of Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5, PDB code: 3cqr
was solved by
P.Arnoux,
T.Morosinotto,
D.Pignol,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 29.20 / 2.00 |
| Space group | I 41 2 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 122.297, 122.297, 158.337, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 20.8 / 23.4 |
Gadolinium Binding Sites:
The binding sites of Gadolinium atom in the Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5
(pdb code 3cqr). This binding sites where shown within
5.0 Angstroms radius around Gadolinium atom.
In total 3 binding sites of Gadolinium where determined in the Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5, PDB code: 3cqr:
Jump to Gadolinium binding site number: 1; 2; 3;
In total 3 binding sites of Gadolinium where determined in the Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5, PDB code: 3cqr:
Jump to Gadolinium binding site number: 1; 2; 3;
Gadolinium binding site 1 out of 3 in 3cqr
Go back to
Gadolinium binding site 1 out
of 3 in the Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5
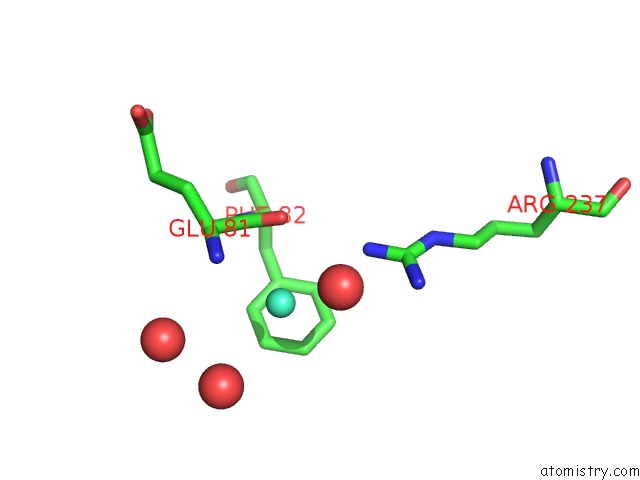
Mono view
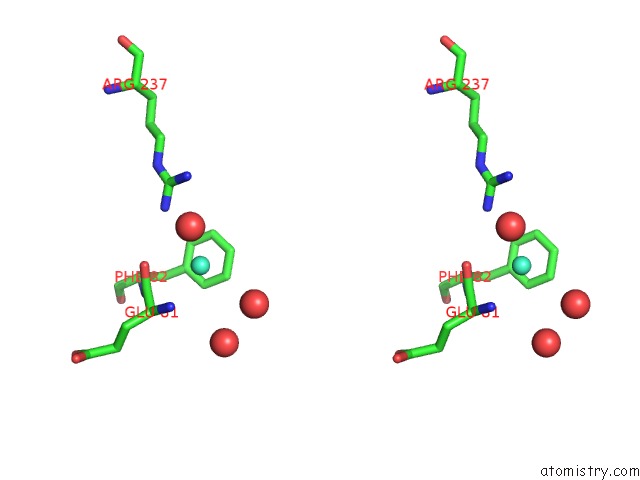
Stereo pair view

Mono view

Stereo pair view
A full contact list of Gadolinium with other atoms in the Gd binding
site number 1 of Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5 within 5.0Å range:
|
Gadolinium binding site 2 out of 3 in 3cqr
Go back to
Gadolinium binding site 2 out
of 3 in the Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5
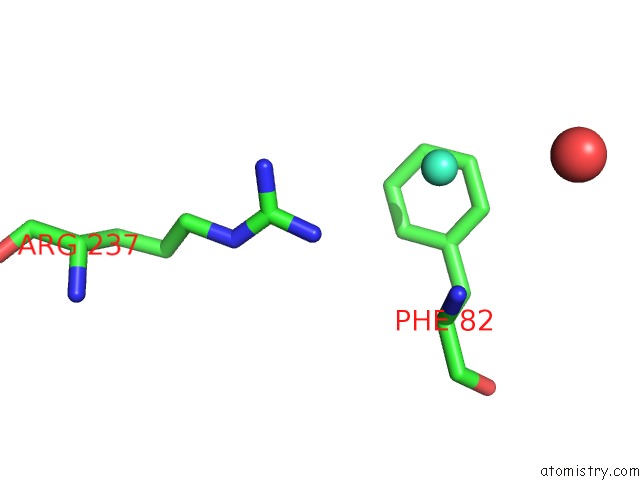
Mono view
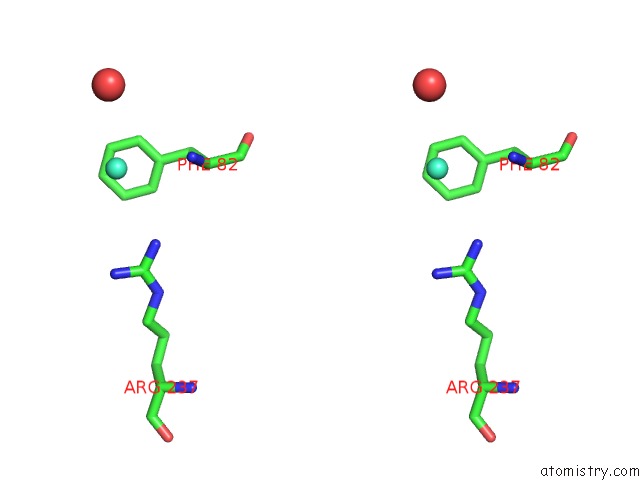
Stereo pair view

Mono view

Stereo pair view
A full contact list of Gadolinium with other atoms in the Gd binding
site number 2 of Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5 within 5.0Å range:
|
Gadolinium binding site 3 out of 3 in 3cqr
Go back to
Gadolinium binding site 3 out
of 3 in the Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5
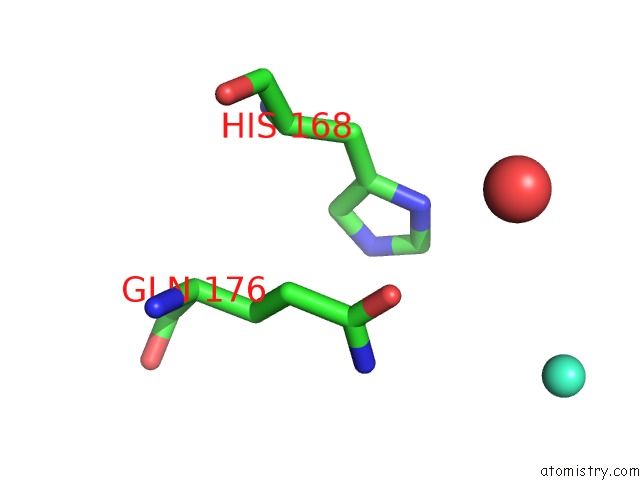
Mono view
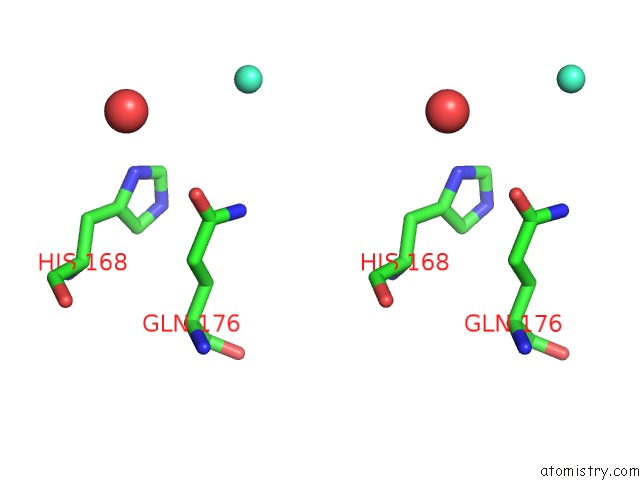
Stereo pair view

Mono view

Stereo pair view
A full contact list of Gadolinium with other atoms in the Gd binding
site number 3 of Crystal Structure of the Lipocalin Domain of Violaxanthin De-Epoxidase (Vde) at PH5 within 5.0Å range:
|
Reference:
P.Arnoux,
T.Morosinotto,
G.Saga,
R.Bassi,
D.Pignol.
A Structural Basis For the pH-Dependent Xanthophyll Cycle in Arabidopsis Thaliana. Plant Cell V. 21 2036 2009.
ISSN: ISSN 1040-4651
PubMed: 19638474
DOI: 10.1105/TPC.109.068007
Page generated: Sat Aug 10 21:23:03 2024
ISSN: ISSN 1040-4651
PubMed: 19638474
DOI: 10.1105/TPC.109.068007
Last articles
As in 6FMIAs in 6EU7
As in 6EXU
As in 6EZV
As in 6EB1
As in 6EB2
As in 6E66
As in 6DUN
As in 6D8P
As in 6CZ8